OverExpress Electrocompetent Cells
OverExpress C41(DE3) Electrocompetent Cells, 12 Reactions (SOLOs)
Express difficult or toxic proteins using standard E. coli T7
expression vectors using these unique strains. Also available in
chemically competent format
Each OverExpress kit contains Electrocompetent Cells in SOLO packaging
(1 transformation per tube), Expression Recovery Medium (lactose minus),
pUC19 Positive Control Plasmid, pAVD10 Verification Plasmid, and
complete protocols.
Product information
E. coli BL21(DE3) strains, like E. cloni™ EXPRESS
Competent Cells provide reliable expression of many genes cloned into
T7 expression vectors (e.g.., pET or pSMART™-cDNA vectors). However, in
some cases expression is minimal or not detectable because the
recombinant protein, when expressed, is deleterious or lethal to these
standard BL21 strains. Examples of such toxic proteins include many
membrane proteins, some cytoplasmic proteins, and nucleases.
Unfortunately, successful expression of one or more toxic proteins is
often important to the experimental goal.
OverExpress Electrocompetent and Chemically Competent Cells are E. coli strains
that are effective in expressing toxic proteins from all classes of
organisms, including eubacteria, yeasts, plants, viruses, and mammals.
The effectiveness of these new strains in expressing toxic proteins has
been validated in more than 350 publications.
The OverExpress strains contain genetic mutations phenotypically
selected for conferring tolerance to toxic proteins. The strain C41(DE3)
was derived from BL21(DE3). This strain has at least one mutation,
which prevents cell death associated with expression of many recombinant
toxic proteins. The strain C43(DE3) was derived from C41(DE3) by
selecting for resistance to a different toxic protein and can express a
different set of toxic proteins to C41(DE3). Figure. 1 graphically
illustrates the advantages of the OverExpress Competent Cells, compared
to standard BL21(DE3) cells, in expressing toxic proteins.
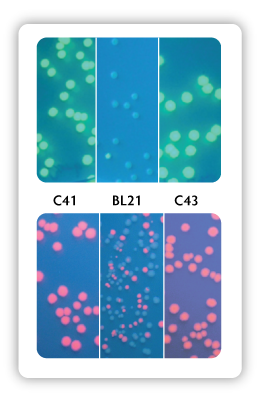
Figure 1. Green Fluorescent Protein (top) or Red
Fluorescent Protein (bottom) expressed from a T7 promoter construct that
was transformed into C41, BL21, or C43 competent cells spread on IPTG
plates to induce protein expression.
Table. 1 and Figure. 2 summarise transformation effectiveness,
tolerance of expression-induced toxicity, and protein expression for T7
expression plasmids coding for a variety of recombinant proteins. These
results demonstrate that the OverExpress C41(DE3) and C43(DE3) strains
are clearly superior to the parental BL21(DE3) in transformation and
expression of toxic proteins.
Table 1. Comparison of OverExpress C41(DE3) and
C43(DE3) cells with the parental strain BL21(DE3) in transformation and
expression of heterologous proteins.**
| Strain |
Transformation
Success Ratea
|
Expression-induced Toxicityb
|
Expressing Plasmidsc
|
|
BL21(DE3)
|
16/26 (62%)
|
25/26 (96%)
|
14/26 (54%)
|
|
C41(DE3)
|
28/28 (100%)
|
14/28 (50%)
|
24/28 (86%)
|
|
C43(DE3)
|
28/28 (100%)
|
1/28 (4%)
|
23/28 (81%) |
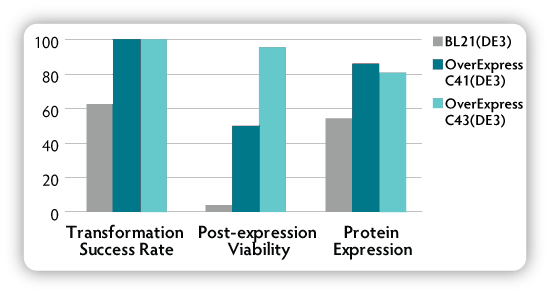
Figure 2. Comparison of OverExpress C41(DE3) and
C43(DE3) cells with the parental strain BL21(DE3) in transformation and
expression of heterologous proteins.**
a Transformation success corresponds to the presence of colonies on LB ampicillin agar following transformation with a plasmid.
b Expression toxicity corresponds to the absence of colonies on LB ampicillin IPTG agar following transformation with a plasmid.
c
Expressing plasmids corresponds to observation of a heterologous
protein in the total cell pellet on Coomassie-stained SDS-PAGE following
growth of a colony in LB ampicillin medium and induction with IPTG.
**L. Dumon-Seignovert, G. Cariot, and L. Vuillard (2004). Protein Expression and Purification 37, 203-206. Data used with permission.
As in standard BL21(DE3) strains, OverExpress C41(DE3),
C41(DE3)pLysS, C43(DE3), and C43 (DE3)pLysS are lysogens of λDE3. These
strains carry a chromosomal copy of the T7 RNA Polymerase gene under the
control of the lacUV5 promoter. These strains are suitable for
production of protein from target genes cloned into T7-driven
expression vectors. OverExpress C41(DE3), C41(DE3) pLysS, C43(DE3), and
C43(DE3) pLysS are also deficient in the lon and ompT proteases.
OverExpress C41(DE3)pLysS and C43(DE3)pLysS also carry a
chloramphenicol-resistant plasmid that encodes T7 lysozyme, which is a
natural inhibitor of T7 RNA polymerase. Cells containing pLysS produce a
small amount of T7 lysozyme. These strains are used to suppress basal
expression of T7 RNA polymerase prior to induction, thus stabilising
recombinants encoding particularly toxic proteins.
OverExpress Genotypes
OverExpress C41(DE3): F – ompT hsdSB (rB- mB-) gal dcm (DE3)
OverExpress C41(DE3)pLysS: F – ompT hsdSB (rB- mB-) gal dcm (DE3) pLysS (CmR)
OverExpress C43(DE3): F – ompT hsdSB (rB- mB-) gal dcm (DE3)
OverExpress C43(DE3)pLysS: F – ompT hsdSB (rB- mB-) gal dcm (DE3) pLysS (CmR)
Table 1. OverExpress Transformation Efficiencies
| Electrocompetent Cells |
cfu/µg DNA |
| C41(DE3) |
> 1 × 109 |
| C43(DE3) |
> 1 × 109 |
| Chemically Competent Cells |
| C41(DE3) |
> 1 × 106 |
| C41(DE3)pLysS |
> 1 × 106 |
| C43(DE3) |
> 1 × 106 |
| C43(DE3)pLysS |
> 1 × 106 |
SDS
Manuals and user guides
Product information sheets
If you cannot find the answer to your problem then please contact us or telephone +44 (0)1954 210 200