Transposomics
The field of Transposomics exploits the ability of certain transposase enzymes to catalyze the random hop or insertion of an “artificial” transposon into any other DNA during a brief in vitro or in vivo transposition reaction. This range of products utilise simple in vitro and in vivo reactions which are fast, reliable and do not require previous knowledge of transposomics. Our EZ::Tn5 in vitro transposition systems can be used to simplify and speed up DNA sequencing. In Vivo Transposomics is revolutionizing microbial genetics by providing tools for studying organisms for which genetic tools were not previously available. These new transposon tools are ideal for protein engineering and mapping of protein domains or epitopes.
The Applications of Transposomics™ Are Limited Only by Our Imagination
The field of Transposomics™ exploits the ability of certain transposase enzymes to catalyze the random “hop” or insertion of an “artificial” transposon into any other DNA during a brief in vitro or in vivo transposition reaction. The transposon can be any DNA that has a transposase recognition sequence on each end (e.g., Figure 1). An artificial transposon doesn't encode a transposase, but can encode selectable markers, priming sites, origins of replication, RNA polymerase promoters, or other genetic elements or control sequences.
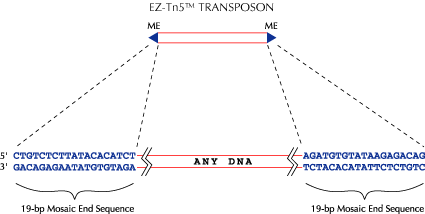
Figure 1.
An EZ-Tn5™ Transposon. An EZ-Tn5 Transposon can be any DNA sequence that is between properly-oriented 19-bp inverted repeat Mosaic End (ME) sequences that are specifically and uniquely recognized by EZ-Tn5™ Transposase. The EZ-Tn5 System is based on the hyperactive Tn5 in vitro transposition system described by IY Goryshin & WS Reznikoff1. This system retains the insertion characteristics of Tn5–the most random transposon system known–but has a transposition frequency 1000-fold higher than wild-type Tn5. EPICENTRE also has a HyperMu™ In Vitro Transposition System that is 50-100 times more active than other MuA Transposase Systems
.
Webinar
How can Transposmic accelerate you genomic research